Please acknowledge the MRCF in your publications:
“Idaho State University Molecular Research Core Facility, RRID:SCR_012598”
We’d like to keep track of our scientific contributions. If we have contributed in any way with your research, then please acknowledge us when that research is published. If possible, please send us an email when your research is published. For help writing Materials and Methods of services provided, please email the lab - we can provide information about our protocols and instruments used for your research project.
Services Provided by the MRCF
The MRCF offers a wide variety of molecular services. We accept whole tissues all the way through library prepped samples, and are willing to assist at any step of the way for preparing your samples for analysis. We are always willing to help, and if you are looking for a service that you don't see listed, let us know and we will discuss the option of optimizing and adding it to our list of services we can provide.
The cost of our services can vary depending on the current cost of reagents. We do have a price list prepared but for the most accurate pricing please email us at mrcf@isu.edu.
Next-Generation DNA Sequencing
Impact:
Next-Generation Sequencing (NGS) allows for the characterization of unculturable microorganisms, the discovery of new viruses, and provides insight into new outbreak control methods in the field of microbial genomics.
Here is a list of just a few of the applications NGS is used for:
Agrigenomics
Oncology
Forensic Genomics
Complex Disease Genomics
Microbial Genomics and Environmental Profiling
Genomics in Drug Development
Reproductive and Genetic Health
And More...

NGS has revolutionized how genomes and DNA sequencing can further scientific research. Read more at Illumina's website for project design ideas and inspiration.
The MRCF provides library preparation for most types of NGS projects on Illumina instruments, as well as sequencing.
Sanger - DNA Sequencing
Sanger sequencing is performed on our Seq Studio Flex 8-capillary genetic analyzer. When sequencing we will include at least one positive control.
Please submit an MRCF Job Request Form with your samples. This is a downloadable Excel file that you will fill out and email an electronic copy to mrcf@isu.edu, as well as send a hard copy along with the samples submitted/shipped to us.
First time customers, will also need to submit a New User Form.
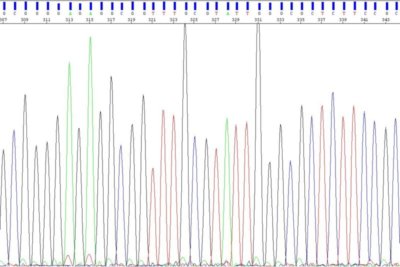
Fragment Analysis
Fragment Analysis on Sanger Sequencer - Microsatellites and T-RFLPs
Microsatellite markers, also known as short tandem repeats (STRs), are polymorphic DNA loci consisting of a repeated nucleotide sequence. The number of repeats varies in a population, thus creating multiple alleles for a microsatellite locus. Microsatellite loci are amplified by PCR using fluorescently labeled forward primer. These PCR amplicons are then separated by size on the Seq Studio Flex genetic analyzer. T-RFLP analysis is a technique used to study complex microbial communities based on variation in the 16S rRNA gene. It can be used to study microbial community structure, bacterial populations in the environment and changes in a microbial community in response to different environmental parameters.
Our instrument runs batches of 8 samples - but entire plates can be set-up and scheduled to run on the instrument.
Available Size Standards: LIZ600 & ROX1000
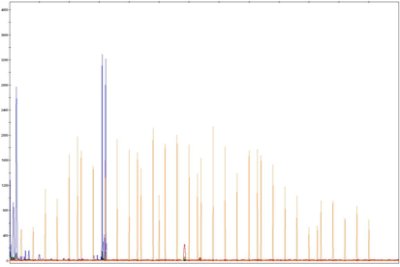
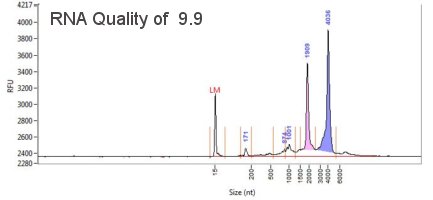
Agilent Fragment Analyzer - DNA/RNA Profiling
The Fragment Analyzer covers the full spectrum of DNA, gDNA and RNA quality control. It is used and recommended by Illumina as an automated system for the quantification and qualification of NGS libraries, gDNA and RNA. Our instrument has 12-capillaries that electrophorese 11-fluorescently labeled samples and a size standard (ladder) per instrument run. It can resolve fragments from 10-40,000 bp in less than 1 hour. The instrument has a resolution of 2bp for fragments which can be detected in as little concentration as 5 pg/uL. The instrument is one of the best to look at the profile, integrity and quality of samples.
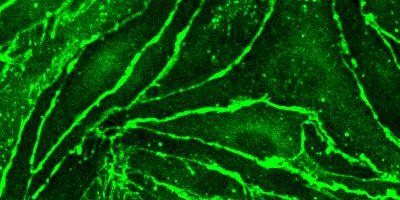
Microscopy and Histology
Olympus FV1000MPE Multiphoton (2P) Microscope
The excitation from the multiphoton microscope is superior for imaging living cells that reside within intact tissues such as brain slices, embryos, whole organs, and even entire animals. The technique provides optical sectioning with reduced absorption of excitation light in specimen regions removed from the objective focal plane, thus minimizing photobleaching and phototoxicity. Please talk to MRCF staff to see if this system is right for you!
Olympus FV1000 Confocal Microscope
Our confocal laser scanning microscope has four simultaneous detection channels, including one for DIC and two spectral channels. The lasers used for excitation on our instrument cover the visible spectrum; the exact wavelengths are 405nm, 458nm, 488nm, 515nm, 559nm and 635nm. It is important to talk to one of our staff members before setting up an experiment to make sure there won't be crosstalk or crossing of two spectrally-similar colors.
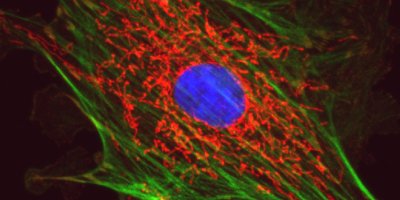
Leica DM6B Upright Widefield Microscope
The DM6B is a fully automated microscope capable of brightfield, darkfield, DIC, phase contrast, and epifluorescence imaging. This scope can also simultaneously capture fluorescence and DIC, and fluorescence and phase images. The ease of use makes this microscope ideal for teaching courses, and great for quick sample screening.
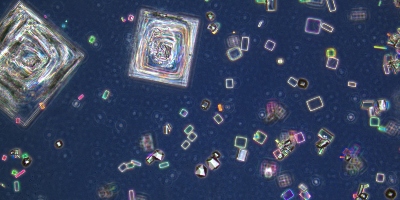

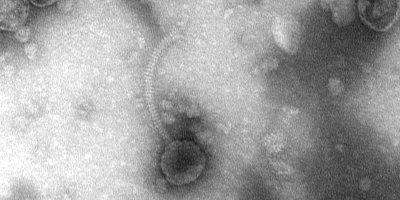
Zeiss EM900 Transmission Electron Microscope
The Transmission Electron Microscope (TEM) is capable of imaging ultra-thin specimen at the magnification of 140,000X, - easily detecting targets as small as cellular organelles and viruses. Use of the TEM requires meticulous sample preparation of tissue support mesh grids coated in a layer of carbon molecules. Our Gatan CCD digital camera makes image capture quick and easy. Sample preparation is specialized for this instrument, please contact MRCF staff with questions about this process.
Leica 3050 S Cryostat
When working with delicate specimens, the precise specimen orientation and the specimen feed system of the Leica cryostat guarantees reproducible, thin, serial sections of high quality. The 3050 S is capable of automatic sectioning with a foot-pedal drive for user convenience.

Leica RM2255 Microtome
Designed for fully motorized sectioning of both paraffin and resin embedded specimens, the Leica RM2255 supports a broad spectrum of applications.

Offline Data Analysis Station
This offline workstation includes multiple programs to help researchers analyze data generated from the Advanced Imaging Facility, and produce publication worthy images and pictures for various uses. Programs available on this computer are MetaMorph for Olympus, ImageJ (Fiji), Vaa3D - 3D Visualization-Assisted Analysis, velocity Demo 6.1.1 and FluoRender. In addition to analysis software, the computer includes Microsoft Office and Adobe Reader.
Flow Cytometry
Flow Cytometry is the analysis of fluorescent markers and dyes on virtually any cell or cell-sized particle. It is a simple, rapid and powerful tool that takes advantage of fluidics, laser and PMT technology, and high-tech electronics for detection and analysis of fluorescence.
The FACSMelody incorporates efficient optics and three solid-state lasers; violet, blue and red; to analyze up to nine colors. The FACSMelody is a highly sensitive tool, providing researchers with the technological capabilities crucial for sophisticated cellular and molecular work in areas such as cell cycle, cell function, multicolor analysis, and immune function.
Our instrument has a sorter that allows researchers to sort out populations of interest, and save them for other applications. The sorter is able to sort at around 34,000 drops/second, which is fast enough to collect large numbers of cells in a fairly short time, and keep them viable and ready to use. It has the ability to sort into a variety of container types: tubes, plates and microscope slides.
Other Services
The MRCF is provides a wide diversity of applications, all at one core facility! If there is a service you require, that isn't listed below, please contact us - we are more than happy to develop and optimize new techniques and when feasible, offer new services.
ELISA Immunosorbent Assays
DNA Extractions
RNA Extractions
qPCR and PCR
Lab Information
Map | Map to MRCF
Hours of Operation | 8:00AM - 4:00PM; Excluding University closures and holidays

